A scientific and personal journey from just 145 base-pairs to thousands of newly diagnosed patients
Written by Nicky Whiffin
In the last two years my science world has been turned completely upside down. The starting point: a discovery in early 2024 of a highly prevalent yet previously unrecognised neurodevelopmental disorder caused by variants in a teeny tiny gene, RNU4-2 (1, 2).
At just 145 DNA bases long, RNU4-2 stands in stark contrast to the massive protein-coding genes that are known to be major players in brain development, and in which DNA variants lead to disease. RNU4-2 is 100-times smaller than most of those genes. And yet RNU4-2 more than pulls its weight: variants in the centre of the gene (in a contiguous region of just 18 bases) cause a highly prevalent neurodevelopmental disorder, ReNU syndrome, which is predicted to impact ~100,000 individuals worldwide.
Just by itself, this discovery was striking: why did we not already know about a disorder so prevalent? Most disorders of comparable frequency were discovered by the early 2010s, when we first had access to large exome sequenced patient cohorts. There are perhaps two key contributing factors: (1) RNU4-2 is a non-coding gene, with the vast majority of our gene discovery efforts to date focussed solely on genes that encode proteins; (2) there isn’t an obvious ‘phenotypic hook’ or super specific symptom that would have prompted targeted investigation into a subset of ReNU patients - although some dysmorphologists think that we perhaps should have grouped ReNU patients based on their distinctive facial features.
But what amazes me the most is what has happened since those first two papers were published. Two things have become abundantly clear to me: (1) science can move at an incredible pace of discovery, especially when given a new and exciting focus and when it is grounded in teamwork and collaboration; and (2) families affected by genetic disorders have incredible drive, focus, and strength - they can quickly build an engaged and supportive community, and joining that community as a scientist is hugely rewarding.
RNUopathies: a rapidly growing class of disorders caused by variants in RNU genes
The RNU4-2 gene is the instructions to make a small nuclear RNA (snRNA) called U4. U4 is one of nine snRNAs that are all critical RNA components of the ‘spliceosome’ complexes that mediate RNA splicing (there are two, the ‘major’ and the ‘minor’). There are hundreds of genes in the human genome that produce these snRNAs (along with many pseudogenes) the names of which all start with the same three letters: ‘RNU’.
RNU4-2 was not the first snRNA gene to be associated with a rare disorder; that crown belongs to RNU4ATAC, which was first linked to disease back in 2011 and is now associated with three named disorders (3). Variants in RNU12 were also linked to two rare disorders in 2017 and 2021 (4). But 2024 was perhaps the first time that many people properly paused to consider a broader role for these small RNA genes. The discovery of ReNU syndrome came at a time when large genome sequencing cohorts exist around the world, enabling people to look at these tiny non-coding genes, but also, ReNU syndrome is far more common than the rare disorders caused by variants in RNU4ATAC and RNU12.
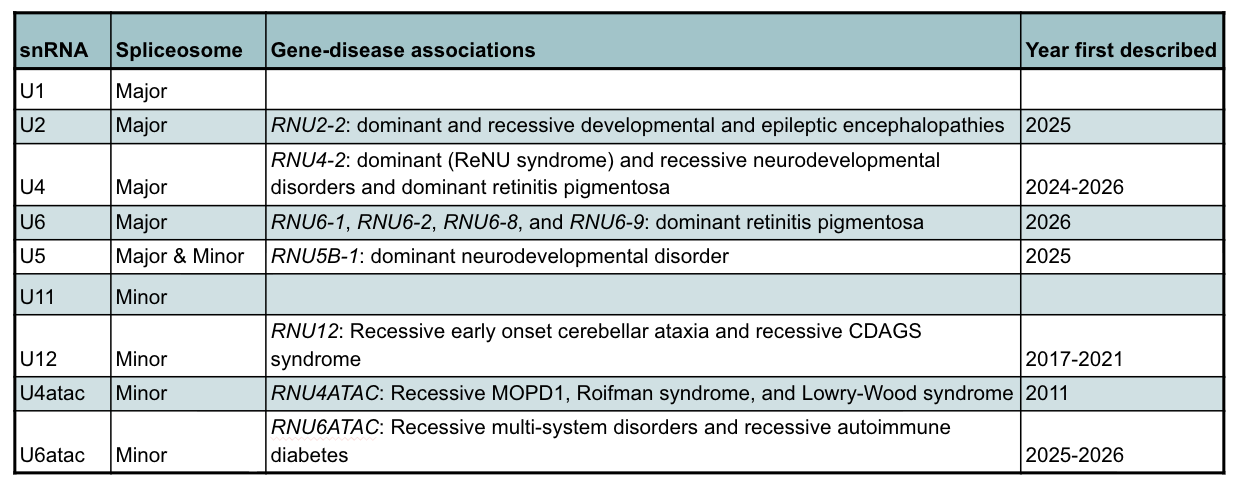
At the time of writing, there are now 17 distinct associations between rare genetic disorders and snRNA genes (Table) that can collectively be referred to as ‘RNUopathies’. Variants in RNU2-2 and RNU5B-1 were, in 2025, linked to dominant neurodevelopmental disorders (5, 6, 7), and two weeks ago, three papers described the discovery of a highly prevalent recessive disorder, also associated with RNU2-2 (8, 9, 10). Variants in RNU4-2 and four genes that produce the U6 snRNA were earlier this year associated with retinitis pigmentosa (11), and just last week, two papers were published describing our discovery of a recessive neurodevelopmental disorder associated with RNU4-2 (12, 13). Finally, in the minor spliceosome, three recent papers describe variants in RNU6ATAC in developmental disorders (14, 15) and early-onset autoimmune diabetes (16).
That is twelve new disorders associated with eight snRNA genes discovered within the space of two years.
Collectively, these disorders likely affect >200,000 individuals globally. The eight genes, they take up just 1,004 bases of genome ‘real estate’ (or 0.00003% of our DNA). But we are still only just beginning to understand the role of variation in these genes and disease risk, and it is truly exciting to see where the next two years of discoveries take us.
Table: RNUopathies, the associated snRNAs, and the year they were first described.
The majority of these recent discoveries have relied on large-scale international collaboration and teamwork both within and between groups. We are at an incredible time for rare disease research whereby large genome datasets exist across the world and sometimes a 5 minute look-up while on a conference call is enough to identify multiple additional patients. It is telling that some of the key papers describing these discoveries have >100 authors (a real pain to enter into the journal submission website!!).
On a personal note, the collaborations, and friendships, that have been built through these projects over these past two years will likely last a lifetime.
The incredible power of advocacy and community
Very soon after receiving her son Devon’s diagnosis of ReNU syndrome in April 2024, Jessica Margrill set up a Facebook group to try to connect with other families. The group has grown dramatically and is now a family support group with over 680 members. Jessica, her sister Heather, and two other ReNU mums, Lindsay and Lauren, have since established the not-for-profit US-based foundation ReNU syndrome United. Similar patient advocacy groups now exist in at least six countries, including the UK, and more recently founded groups focus on the other RNUopathies. There is also an umbrella organisation, covering all of the RNUopathies. The aim of these groups is simple: build community, raise awareness, advocate for genetic testing of snRNA genes, and drive forward research into developing treatments.
ReNU syndrome United has achieved an incredible amount already. You only have to spend five minutes on their website (www.renusyndrome.org/) to realise this. I am repeatedly blown away by their ability to convince researchers, clinicians, and industry partners to join the ReNU community. They are connected to a multitude of research studies, hold regular awareness days and fundraising campaigns, and even have a shop with a vast array of branded swag (that of course I couldn’t resist getting some of). Sometimes it is impossible to believe that all this has been achieved in only two years. What is even more remarkable, is that these incredible parents have managed this alongside full time jobs and while caring for a severely disabled child. They are, quite simply, superheroes.
For me, interacting with the ReNU community has brought me far closer to rare disease families than I have ever been before. Previously, like many academics, I was at least one step removed from direct interactions. But being closer to the families brings an incredible clarity to exactly why we do what we do and is hugely rewarding and inspiring.
In July last year, I had the enormous privilege of attending the inaugural ReNU Hope conference in Long Island, New York (17). It was an impressively professional set-up with over 200 attendees over two days; the first focussed on the families and the second a scientific symposium. This was the first time that I met ReNU warriors and their families in person. There were a lot of tears, however, the predominant feeling was one of immense joy and a sense of community. I even got to dance with the amazing Devon. Smaller scale events have now taken place in other countries, including France and Germany, and I am looking forward to the first UK family meet-up in the UK in a couple of months.
Left to right: (1) Meeting the amazing Jessica. (2) Devon and his sister. (3) ReNU Syndrome United’s Supermoms.
ReNU’d hope for therapies
But the science doesn’t stop with the discovery of these disorders. Rather, this new knowledge gives us an opportunity; while without a genetic diagnosis treatment options for patients are limited to treating only symptoms, knowing the root-cause of a disorder allows us to target it directly.
Recently, advances in genome targeting therapies, including CRISPR and antisense oligonucleotides (ASOs), mean that these RNUopathy discoveries come at a time when we have the tools to potentially treat them. Moreover, ReNU syndrome hits that sweet-spot for a rare disorder where it is common enough to be an attractive target for pharmaceutical companies and for us to be able to conduct clinical trials. In both research and industry settings ASOs have already been identified that can selectively reduce the mutant U4 RNA in cells from individuals with ReNU syndrome. There is still a lot of uncertainty, however, and while scientifically we can develop a molecule with the desired impact, this does not mean that it will improve the features of ReNU syndrome. We still have a very long way to go before we have a treatment that is both safe and effective.
That said, there is an incredible amount of hope for the thousands of families with these disorders that just didn’t exist two years ago.
References
Chen, Y. et al. Nature 632, 832–840 (2024).
Greene, D. et al. Nature Medicine 30, 2165–2169 (2024).
Nava, C. et al. Nature Genetics 57, 1374–1388 (2025).
Greene, D. et al. Nature Genetics 57, 1367–1373 (2025).
Jackson, A. et al. Nature Genetics 57, 1362–1366 (2025).
Leitão, E. et al. Nature Genetics (2026).
Jackson, A. et al. Nature Genetics (2026).
Greene, D. et al. Nature Genetics (2026).
Quinodoz, M. et al. Nature Genetics 58, 169–179 (2026).
De Jonghe, J. et al. Nature (2026).
Rius, R. et al. Nature Genetics (2026).
Arriaga, T.M. et al. AJHG 112, 2458-2475 (2025).
Mendez, R. et al. HGG Advances (2026).
Johnson, M.B. et al. AJHG 113, 877-887 (2026).
Crocker, K., et al. American Journal of Medical Genetics Part A1–7 (2026).